Python API¶
This section includes information for using the pure Python API of bob.learn.em.
Classes¶
Trainers¶
| bob.learn.em.KMeansTrainer | Trains a KMeans machine.This class implements the |
| bob.learn.em.ML_GMMTrainer | This class implements the maximum likelihood M-step of the expectation-maximisation algorithm for a GMM Machine. |
| bob.learn.em.MAP_GMMTrainer | This class implements the maximum a posteriori M-step of the expectation-maximization algorithm for a GMM Machine. |
| bob.learn.em.ISVTrainer | ISVTrainerReferences: [Vogt2008,McCool2013] |
| bob.learn.em.JFATrainer | JFATrainerReferences: [Vogt2008,McCool2013] |
| bob.learn.em.IVectorTrainer | IVectorTrainerAn IVectorTrainer to learn a Total Variability subspace 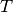 (and eventually a covariance matrix (and eventually a covariance matrix 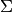 ). ). |
| bob.learn.em.PLDATrainer | This class can be used to train the 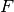 , , 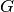 and and |
| bob.learn.em.EMPCATrainer | Constructor Documentation: |
Machines¶
| bob.learn.em.KMeansMachine | This class implements a k-means classifier. |
| bob.learn.em.Gaussian | This class implements a multivariate diagonal Gaussian distribution |
| bob.learn.em.GMMStats | A container for GMM statistics |
| bob.learn.em.GMMMachine | This class implements a multivariate diagonal Gaussian distribution. |
| bob.learn.em.ISVBase | A ISVBase instance can be seen as a container for U and D when performing Joint Factor Analysis (JFA). |
| bob.learn.em.ISVMachine | A ISVMachine. |
| bob.learn.em.JFABase | A JFABase instance can be seen as a container for 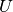 , , 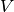 and and 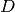 when performing Joint Factor Analysis (JFA). when performing Joint Factor Analysis (JFA). |
| bob.learn.em.JFAMachine | A JFAMachine. |
| bob.learn.em.IVectorMachine | An IVectorMachine consists of a Total Variability subspace 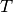 |
| bob.learn.em.PLDABase | This class is a container for the 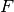 (between class variantion matrix), (between class variantion matrix), 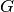 (within class variantion matrix) and (within class variantion matrix) and 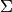 matrices and the mean vector matrices and the mean vector 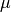 of a PLDA model. of a PLDA model. |
| bob.learn.em.PLDAMachine | This class is a container for an enrolled identity/class. |
Functions¶
| bob.learn.em.linear_scoring((models, ubm, ...) | Parameters: |
| bob.learn.em.tnorm(...) | Normalise raw scores with T-Norm |
| bob.learn.em.train(trainer, machine, data[, ...]) | Trains a machine given a trainer and the proper data |
| bob.learn.em.train_jfa(trainer, jfa_base, data) | Trains a bob.learn.em.JFABase given a bob.learn.em.JFATrainer and the proper data |
| bob.learn.em.znorm(...) | Normalise raw scores with Z-Norm |
| bob.learn.em.ztnorm(...) | Normalise raw scores with ZT-Norm.Assume that znorm and tnorm have no common subject id. |
| bob.learn.em.ztnorm_same_value(vect_a, vect_b) | Computes the matrix of boolean D for the ZT-norm, which indicates where the client ids of the T-Norm models and Z-Norm samples match. |
Detailed Information¶
- bob.learn.em.ztnorm_same_value(vect_a, vect_b)[source]¶
Computes the matrix of boolean D for the ZT-norm, which indicates where the client ids of the T-Norm models and Z-Norm samples match.
vect_a An (ordered) list of client_id corresponding to the T-Norm models vect_b An (ordered) list of client_id corresponding to the Z-Norm impostor samples
- class bob.learn.em.EMPCATrainer¶
Bases: object
Constructor Documentation:
- bob.learn.em.EMPCATrainer (convergence_threshold)
- bob.learn.em.EMPCATrainer (other)
- bob.learn.em.EMPCATrainer ()
Creates a EMPCATrainer
Parameters:
other : bob.learn.em.EMPCATrainer
A EMPCATrainer object to be copied.convergence_threshold : float
Class Members:
- compute_likelihood(linear_machine, data) → None¶
Todo
The parameter(s) ‘data’ are used, but not documented.
Parameters:
linear_machine : bob.learn.linear.Machine
LinearMachine Object
- e_step(linear_machine, data) → None¶
Parameters:
linear_machine : bob.learn.linear.Machine
LinearMachine Objectdata : array_like <float, 2D>
Input data
- initialize(linear_machine, data) → None¶
Todo
The parameter(s) ‘rng’ are documented, but nowhere used.
Parameters:
linear_machine : bob.learn.linear.Machine
LinearMachine Objectdata : array_like <float, 2D>
Input datarng : bob.core.random.mt19937
The Mersenne Twister mt19937 random generator used for the initialization of subspaces/arrays before the EM loop.
- m_step(linear_machine, data) → None¶
Parameters:
linear_machine : bob.learn.linear.Machine
LinearMachine Objectdata : array_like <float, 2D>
Input data
- class bob.learn.em.GMMMachine¶
Bases: object
This class implements a multivariate diagonal Gaussian distribution.
See Section 2.3.9 of Bishop, “Pattern recognition and machine learning”, 2006
Constructor Documentation:
- bob.learn.em.GMMMachine (n_gaussians,n_inputs)
- bob.learn.em.GMMMachine (other)
- bob.learn.em.GMMMachine (hdf5)
- bob.learn.em.GMMMachine ()
Creates a GMMMachine
Parameters:
n_gaussians : int
Number of gaussiansn_inputs : int
Dimension of the feature vectorother : bob.learn.em.GMMMachine
A GMMMachine object to be copied.hdf5 : bob.io.base.HDF5File
An HDF5 file open for readingClass Members:
- acc_statistics(input, stats) → None¶
Accumulate the GMM statistics for this sample(s). Inputs are checked.
Parameters:
input : array_like <float, 2D>
Input vectorstats : bob.learn.em.GMMStats
Statistics of the GMM
- acc_statistics_(input, stats) → None¶
Accumulate the GMM statistics for this sample(s). Inputs are NOT checked.
Parameters:
input : array_like <float, 2D>
Input vectorstats : bob.learn.em.GMMStats
Statistics of the GMM
- get_gaussian(i) → gaussian¶
Get the specified Gaussian component.
Note
An exception is thrown if i is out of range.
Parameters:
i : int
Index of the gaussianReturns:
gaussian : bob.learn.em.Gaussian
Gaussian object
- is_similar_to(other[, r_epsilon][, a_epsilon]) → output¶
Compares this GMMMachine with the other one to be approximately the same.
The optional values r_epsilon and a_epsilon refer to the relative and absolute precision for the weights, biases and any other values internal to this machine.
Parameters:
other : bob.learn.em.GMMMachine
A GMMMachine object to be compared.r_epsilon : float
Relative precision.a_epsilon : float
Absolute precision.Returns:
output : bool
True if it is similar, otherwise false.
- load(hdf5) → None¶
Load the configuration of the GMMMachine to a given HDF5 file
Parameters:
hdf5 : bob.io.base.HDF5File
An HDF5 file open for reading
- log_likelihood(input) → output¶
Output the log likelihood of the sample, x, i.e.
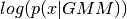 . Inputs are checked.
. Inputs are checked.Note
The __call__() function is an alias for this.
Parameters:
input : array_like <float, 1D>
Input vectorReturns:
output : float
The log likelihood
- log_likelihood_(input) → output¶
Output the log likelihood of the sample, x, i.e.
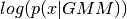 . Inputs are NOT checked.
. Inputs are NOT checked.Parameters:
input : array_like <float, 1D>
Input vectorReturns:
output : float
The log likelihood
- mean_supervector¶
array_like <float, 1D> <– The mean supervector of the GMMMachine
Concatenation of the mean vectors of each Gaussian of the GMMMachine
- means¶
array_like <float, 2D> <– The means of the gaussians
- resize(n_gaussians, n_inputs) → None¶
Allocates space for the statistics and resets to zero.
Parameters:
n_gaussians : int
Number of gaussiansn_inputs : int
Dimensionality of the feature vector
- save(hdf5) → None¶
Save the configuration of the GMMMachine to a given HDF5 file
Parameters:
hdf5 : bob.io.base.HDF5File
An HDF5 file open for writing
- set_variance_thresholds(input) → None¶
Set the variance flooring thresholds in each dimension to the same vector for all Gaussian components if the argument is a 1D numpy arrray, and equal for all Gaussian components and dimensions if the parameter is a scalar.
Parameters:
input : array_like <float, 1D>
Input vector
- shape¶
(int,int) <– A tuple that represents the number of gaussians and dimensionality of each Gaussian (n_gaussians, dim).
- variance_supervector¶
array_like <float, 1D> <– The variance supervector of the GMMMachine
Concatenation of the variance vectors of each Gaussian of the GMMMachine
- variance_thresholds¶
array_like <float, 2D> <– Set the variance flooring thresholds in each dimension to the same vector for all Gaussian components if the argument is a 1D numpy arrray, and equal for all Gaussian components and dimensions if the parameter is a scalar.
- variances¶
array_like <float, 2D> <– Variances of the gaussians
- weights¶
array_like <float, 1D> <– The weights (also known as “mixing coefficients”)
- class bob.learn.em.GMMStats¶
Bases: object
A container for GMM statistics
With respect to [Reynolds2000] the class computes:
- Eq (8) is bob.learn.em.GMMStats.n:
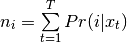
- Eq (9) is bob.learn.em.GMMStats.sum_px:
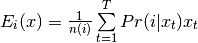
- Eq (10) is bob.learn.em.GMMStats.sum_pxx:
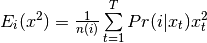
Constructor Documentation:
- bob.learn.em.GMMStats (n_gaussians,n_inputs)
- bob.learn.em.GMMStats (other)
- bob.learn.em.GMMStats (hdf5)
- bob.learn.em.GMMStats ()
A container for GMM statistics.
Parameters:
n_gaussians : int
Number of gaussiansn_inputs : int
Dimension of the feature vectorother : bob.learn.em.GMMStats
A GMMStats object to be copied.hdf5 : bob.io.base.HDF5File
An HDF5 file open for readingClass Members:
- init() → None¶
Resets statistics to zero.
- is_similar_to(other[, r_epsilon][, a_epsilon]) → output¶
Compares this GMMStats with the other one to be approximately the same.
The optional values r_epsilon and a_epsilon refer to the relative and absolute precision for the weights, biases and any other values internal to this machine.
Parameters:
other : bob.learn.em.GMMStats
A GMMStats object to be compared.r_epsilon : float
Relative precision.a_epsilon : float
Absolute precision.Returns:
output : bool
True if it is similar, otherwise false.
- load(hdf5) → None¶
Load the configuration of the GMMStats to a given HDF5 file
Parameters:
hdf5 : bob.io.base.HDF5File
An HDF5 file open for reading
- log_likelihood¶
float <– The accumulated log likelihood of all samples
- n¶
array_like <float, 1D> <– For each Gaussian, the accumulated sum of responsibilities, i.e. the sum of
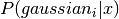
- resize(n_gaussians, n_inputs) → None¶
Allocates space for the statistics and resets to zero.
Parameters:
n_gaussians : int
Number of gaussiansn_inputs : int
Dimensionality of the feature vector
- save(hdf5) → None¶
Save the configuration of the GMMStats to a given HDF5 file
Parameters:
hdf5 : bob.io.base.HDF5File
An HDF5 file open for writing
- shape¶
(int,int) <– A tuple that represents the number of gaussians and dimensionality of each Gaussian (n_gaussians, dim).
- sum_px¶
array_like <float, 2D> <– For each Gaussian, the accumulated sum of responsibility times the sample
- sum_pxx¶
array_like <float, 2D> <– For each Gaussian, the accumulated sum of responsibility times the sample squared
- t¶
int <– The number of samples
- Eq (8) is bob.learn.em.GMMStats.n:
- class bob.learn.em.Gaussian¶
Bases: object
This class implements a multivariate diagonal Gaussian distribution
Constructor Documentation:
- bob.learn.em.Gaussian (n_inputs)
- bob.learn.em.Gaussian (other)
- bob.learn.em.Gaussian (hdf5)
- bob.learn.em.Gaussian ()
Constructs a new multivariate gaussian object
Parameters:
n_inputs : int
Dimension of the feature vectorother : bob.learn.em.GMMStats
A GMMStats object to be copied.hdf5 : bob.io.base.HDF5File
An HDF5 file open for readingClass Members:
- is_similar_to(other[, r_epsilon][, a_epsilon]) → output¶
Compares this Gaussian with the other one to be approximately the same.
The optional values r_epsilon and a_epsilon refer to the relative and absolute precision for the weights, biases and any other values internal to this machine.
Parameters:
other : bob.learn.em.Gaussian
A gaussian to be compared.[r_epsilon] : float
Relative precision.[a_epsilon] : float
Absolute precision.Returns:
output : bool
True if it is similar, otherwise false.
- load(hdf5) → None¶
Load the configuration of the Gassian Machine to a given HDF5 file
Parameters:
hdf5 : bob.io.base.HDF5File
An HDF5 file open for reading
- log_likelihood(input) → output¶
Output the log likelihood of the sample, x. The input size is checked.
Note
The __call__() function is an alias for this.
Parameters:
input : array_like <float, 1D>
Input vectorReturns:
output : float
The log likelihood
- log_likelihood_(input) → output¶
Output the log likelihood given a sample. The input size is NOT checked.
Parameters:
input : array_like <float, 1D>
Input vectorReturns:
output : float
The log likelihood
- mean¶
array_like <float, 1D> <– Mean of the Gaussian
- resize(input) → None¶
Set the input dimensionality, reset the mean to zero and the variance to one.
Parameters:
input : int
Dimensionality of the feature vector
- save(hdf5) → None¶
Save the configuration of the Gassian Machine to a given HDF5 file
Parameters:
hdf5 : bob.io.base.HDF5File
An HDF5 file open for writing
- set_variance_thresholds(input) → None¶
Set the variance flooring thresholds equal to the given threshold for all the dimensions.
Parameters:
input : float
Threshold
- shape¶
(int) <– A tuple that represents the dimensionality of the Gaussian (dim,).
- variance¶
array_like <float, 1D> <– Variance of the Gaussian
- variance_thresholds¶
array_like <float, 1D> <– The variance flooring thresholds, i.e. the minimum allowed value of variance in each dimension.
The variance will be set to this value if an attempt is made to set it to a smaller value.
- class bob.learn.em.ISVBase¶
Bases: object
A ISVBase instance can be seen as a container for U and D when performing Joint Factor Analysis (JFA).
References: [Vogt2008] [McCool2013]
Constructor Documentation:
- bob.learn.em.ISVBase (ubm,ru)
- bob.learn.em.ISVBase (other)
- bob.learn.em.ISVBase (hdf5)
- bob.learn.em.ISVBase ()
Creates a ISVBase
Parameters:
ubm : bob.learn.em.GMMMachine
The Universal Background Model.ru : int
Size of U (Within client variation matrix). In the end the U matrix will have (number_of_gaussians * feature_dimension x ru)other : bob.learn.em.ISVBase
A ISVBase object to be copied.hdf5 : bob.io.base.HDF5File
An HDF5 file open for readingClass Members:
- d¶
array_like <float, 1D> <– Returns the diagonal matrix diag(d) (as a 1D vector)
- is_similar_to(other[, r_epsilon][, a_epsilon]) → output¶
Compares this ISVBase with the other one to be approximately the same.
The optional values r_epsilon and a_epsilon refer to the relative and absolute precision for the weights, biases and any other values internal to this machine.
Parameters:
other : bob.learn.em.ISVBase
A ISVBase object to be compared.r_epsilon : float
Relative precision.a_epsilon : float
Absolute precision.Returns:
output : bool
True if it is similar, otherwise false.
- load(hdf5) → None¶
Load the configuration of the ISVBase to a given HDF5 file
Parameters:
hdf5 : bob.io.base.HDF5File
An HDF5 file open for reading
- resize(rU) → None¶
Resets the dimensionality of the subspace U. U is hence uninitialized.
Parameters:
rU : int
Size of U (Within client variation matrix)
- save(hdf5) → None¶
Save the configuration of the ISVBase to a given HDF5 file
Parameters:
hdf5 : bob.io.base.HDF5File
An HDF5 file open for writing
- shape¶
(int,int, int) <– A tuple that represents the number of gaussians, dimensionality of each Gaussian, dimensionality of the rU (within client variability matrix) (#Gaussians, #Inputs, #rU).
- supervector_length¶
int <– Returns the supervector length.NGaussians x NInputs: Number of Gaussian components by the feature dimensionality
@warning An exception is thrown if no Universal Background Model has been set yet.
- u¶
array_like <float, 2D> <– Returns the U matrix (within client variability matrix)
- ubm¶
bob.learn.em.GMMMachine <– Returns the UBM (Universal Background Model
- class bob.learn.em.ISVMachine¶
Bases: object
A ISVMachine. An attached bob.learn.em.ISVBase should be provided for Joint Factor Analysis. The bob.learn.em.ISVMachine carries information about the speaker factors
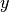 and
and  , whereas a
bob.learn.em.JFABase carries information about the matrices
, whereas a
bob.learn.em.JFABase carries information about the matrices
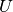 and
and 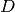 .
.References: [Vogt2008] [McCool2013]
Constructor Documentation:
- bob.learn.em.ISVMachine (isv_base)
- bob.learn.em.ISVMachine (other)
- bob.learn.em.ISVMachine (hdf5)
Constructor. Builds a new ISVMachine
Parameters:
isv_base : bob.learn.em.ISVBase
The ISVBase associated with this machineother : bob.learn.em.ISVMachine
A ISVMachine object to be copied.hdf5 : bob.io.base.HDF5File
An HDF5 file open for readingClass Members:
- estimate_ux(stats, input) → None¶
Estimates Ux (LPT assumption) given GMM statistics.
Estimates Ux from the GMM statistics considering the LPT assumption, that is the latent session variable x is approximated using the UBM.
Parameters:
stats : bob.learn.em.GMMStats
Statistics of the GMMinput : array_like <float, 1D>
Input vector
- estimate_x(stats, input) → None¶
Estimates the session offset x (LPT assumption) given GMM statistics.
Estimates
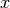 from the GMM statistics considering the LPT
assumption, that is the latent session variable
from the GMM statistics considering the LPT
assumption, that is the latent session variable 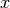 is
approximated using the UBM
is
approximated using the UBMParameters:
stats : bob.learn.em.GMMStats
Statistics of the GMMinput : array_like <float, 1D>
Input vector
- forward_ux(stats, ux) → None¶
Computes a score for the given UBM statistics and given the Ux vector
Parameters:
stats : bob.learn.em.GMMStats
Statistics as inputux : array_like <float, 1D>
Input vector
- is_similar_to(other[, r_epsilon][, a_epsilon]) → output¶
Compares this ISVMachine with the other one to be approximately the same.
The optional values r_epsilon and a_epsilon refer to the relative and absolute precision for the weights, biases and any other values internal to this machine.
Parameters:
other : bob.learn.em.ISVMachine
A ISVMachine object to be compared.r_epsilon : float
Relative precision.a_epsilon : float
Absolute precision.Returns:
output : bool
True if it is similar, otherwise false.
- isv_base¶
bob.learn.em.ISVBase <– The ISVBase attached to this machine
- load(hdf5) → None¶
Load the configuration of the ISVMachine to a given HDF5 file
Parameters:
hdf5 : bob.io.base.HDF5File
An HDF5 file open for reading
- save(hdf5) → None¶
Save the configuration of the ISVMachine to a given HDF5 file
Parameters:
hdf5 : bob.io.base.HDF5File
An HDF5 file open for writing
- shape¶
(int,int, int, int) <– A tuple that represents the number of gaussians, dimensionality of each Gaussian and dimensionality of the rU (within client variability matrix)) (#Gaussians, #Inputs, #rU).
- supervector_length¶
int <– Returns the supervector length.NGaussians x NInputs: Number of Gaussian components by the feature dimensionality
@warning An exception is thrown if no Universal Background Model has been set yet.
- x¶
array_like <float, 1D> <– Returns the
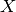 session factor. Eq
(29) from [McCool2013]
session factor. Eq
(29) from [McCool2013]The latent variable x (last one computed). This is a feature provided for convenience, but this attribute is not ‘part’ of the machine. The session latent variable
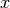 is indeed not class-specific, but
depends on the sample considered. Furthermore, it is not saved into the
machine or used when comparing machines.
is indeed not class-specific, but
depends on the sample considered. Furthermore, it is not saved into the
machine or used when comparing machines.
- z¶
array_like <float, 1D> <– Returns the
 speaker factor. Eq
(31) from [McCool2013]
speaker factor. Eq
(31) from [McCool2013]
- class bob.learn.em.ISVTrainer¶
Bases: object
ISVTrainerReferences: [Vogt2008,McCool2013]
Constructor Documentation:
- bob.learn.em.ISVTrainer (relevance_factor)
- bob.learn.em.ISVTrainer (other)
- bob.learn.em.ISVTrainer ()
Constructor. Builds a new ISVTrainer
Parameters:
other : bob.learn.em.ISVTrainer
A ISVTrainer object to be copied.relevance_factor : float
Class Members:
- acc_u_a1¶
array_like <float, 3D> <– Accumulator updated during the E-step
- acc_u_a2¶
array_like <float, 2D> <– Accumulator updated during the E-step
- e_step(isv_base, stats) → None¶
Call the e-step procedure (for the U subspace).
Parameters:
isv_base : bob.learn.em.ISVBase
ISVBase Objectstats : bob.learn.em.GMMStats
GMMStats Object
- enroll(isv_machine, features, n_iter) → None¶
Parameters:
isv_machine : bob.learn.em.ISVMachine
ISVMachine Objectfeatures : list(bob.learn.em.GMMStats)`
n_iter : int
Number of iterations
- initialize(isv_base, stats, rng) → None¶
Initialization before the EM steps
Parameters:
isv_base : bob.learn.em.ISVBase
ISVBase Objectstats : bob.learn.em.GMMStats
GMMStats Objectrng : bob.core.random.mt19937
The Mersenne Twister mt19937 random generator used for the initialization of subspaces/arrays before the EM loop.
- m_step(isv_base, stats) → None¶
Call the m-step procedure (for the U subspace).
Parameters:
isv_base : bob.learn.em.ISVBase
ISVBase Objectstats : bob.learn.em.GMMStats
Ignored
- class bob.learn.em.IVectorMachine¶
Bases: object
An IVectorMachine consists of a Total Variability subspace
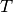 and allows the extraction of IVectorReferences: [Dehak2010]
and allows the extraction of IVectorReferences: [Dehak2010]Constructor Documentation:
- bob.learn.em.IVectorMachine (ubm,rt,variance_threshold)
- bob.learn.em.IVectorMachine (other)
- bob.learn.em.IVectorMachine (hdf5)
Constructor. Builds a new IVectorMachine
Parameters:
ubm : bob.learn.em.GMMMachine
The Universal Background Model.rt : int
Size of the Total Variability matrix (CD x rt).variance_threshold : float
Variance flooring threshold for the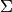 (diagonal)
matrix
(diagonal)
matrixother : bob.learn.em.IVectorMachine
A IVectorMachine object to be copied.hdf5 : bob.io.base.HDF5File
An HDF5 file open for readingClass Members:
- is_similar_to(other[, r_epsilon][, a_epsilon]) → output¶
Compares this IVectorMachine with the other one to be approximately the same.
The optional values r_epsilon and a_epsilon refer to the relative and absolute precision for the weights, biases and any other values internal to this machine.
Parameters:
other : bob.learn.em.IVectorMachine
A IVectorMachine object to be compared.r_epsilon : float
Relative precision.a_epsilon : float
Absolute precision.Returns:
output : bool
True if it is similar, otherwise false.
- load(hdf5) → None¶
Load the configuration of the IVectorMachine to a given HDF5 file
Parameters:
hdf5 : bob.io.base.HDF5File
An HDF5 file open for reading
- project(stats) → None¶
Projects the given GMM statistics into the i-vector subspace
Note
The __call__() function is an alias for this function
Parameters:
stats : bob.learn.em.GMMStats
Statistics as input
- resize(rT) → None¶
Resets the dimensionality of the subspace T.
Parameters:
rT : int
Size of T (Total variability matrix)
- save(hdf5) → None¶
Save the configuration of the IVectorMachine to a given HDF5 file
Parameters:
hdf5 : bob.io.base.HDF5File
An HDF5 file open for writing
- shape¶
(int,int, int) <– A tuple that represents the number of gaussians, dimensionality of each Gaussian, dimensionality of the rT (total variability matrix) (#Gaussians, #Inputs, #rT).
- sigma¶
array_like <float, 1D> <– The residual matrix of the model sigma
- supervector_length¶
int <– Returns the supervector length.NGaussians x NInputs: Number of Gaussian components by the feature dimensionality
@warning An exception is thrown if no Universal Background Model has been set yet.
- t¶
array_like <float, 2D> <– Returns the Total Variability matrix,
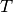
- ubm¶
bob.learn.em.GMMMachine <– Returns the UBM (Universal Background Model)
- variance_threshold¶
float <– Threshold for the variance contained in sigma
- class bob.learn.em.IVectorTrainer¶
Bases: object
IVectorTrainerAn IVectorTrainer to learn a Total Variability subspace
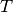 (and eventually a covariance matrix
(and eventually a covariance matrix 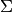 ).References: [Dehak2010]
).References: [Dehak2010]Constructor Documentation:
- bob.learn.em.IVectorTrainer (update_sigma)
- bob.learn.em.IVectorTrainer (other)
- bob.learn.em.IVectorTrainer ()
Constructor. Builds a new IVectorTrainer
Parameters:
other : bob.learn.em.IVectorTrainer
A IVectorTrainer object to be copied.update_sigma : bool
Class Members:
- acc_fnormij_wij¶
array_like <float, 3D> <– Accumulator updated during the E-step
- acc_nij¶
array_like <float, 1D> <– Accumulator updated during the E-step
- acc_nij_wij2¶
array_like <float, 3D> <– Accumulator updated during the E-step
- acc_snormij¶
array_like <float, 2D> <– Accumulator updated during the E-step
- e_step(ivector_machine, stats) → None¶
Call the e-step procedure (for the U subspace).
Parameters:
ivector_machine : bob.learn.em.ISVBase
IVectorMachine Objectstats : bob.learn.em.GMMStats
GMMStats Object
- initialize(ivector_machine[, stats][, rng]) → None¶
Initialization before the EM steps
Parameters:
ivector_machine : bob.learn.em.IVectorMachine
IVectorMachine Objectstats : bob.learn.em.GMMStats
Ignoredrng : bob.core.random.mt19937
The Mersenne Twister mt19937 random generator used for the initialization of subspaces/arrays before the EM loop.
- m_step(ivector_machine, stats) → None¶
Call the m-step procedure (for the U subspace).
Parameters:
ivector_machine : bob.learn.em.ISVBase
IVectorMachine Objectstats : bob.learn.em.GMMStats
Ignored
- class bob.learn.em.JFABase¶
Bases: object
A JFABase instance can be seen as a container for
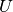 ,
, 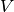 and
and 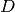 when performing Joint Factor Analysis (JFA).
when performing Joint Factor Analysis (JFA).References: [Vogt2008] [McCool2013]
Constructor Documentation:
- bob.learn.em.JFABase (ubm,ru,rv)
- bob.learn.em.JFABase (other)
- bob.learn.em.JFABase (hdf5)
- bob.learn.em.JFABase ()
Constructor. Builds a new JFABase
Parameters:
ubm : bob.learn.em.GMMMachine
The Universal Background Model.ru : int
Size of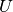 (Within client variation matrix). In the end
the U matrix will have (#gaussians * #feature_dimension x ru)
(Within client variation matrix). In the end
the U matrix will have (#gaussians * #feature_dimension x ru)rv : int
Size of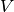 (Between client variation matrix). In the end
the U matrix will have (#gaussians * #feature_dimension x rv)
(Between client variation matrix). In the end
the U matrix will have (#gaussians * #feature_dimension x rv)other : bob.learn.em.JFABase
A JFABase object to be copied.hdf5 : bob.io.base.HDF5File
An HDF5 file open for readingClass Members:
- d¶
array_like <float, 1D> <– Returns the diagonal matrix
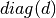 (as a 1D vector)
(as a 1D vector)
- is_similar_to(other[, r_epsilon][, a_epsilon]) → output¶
Compares this JFABase with the other one to be approximately the same.
The optional values r_epsilon and a_epsilon refer to the relative and absolute precision for the weights, biases and any other values internal to this machine.
Parameters:
other : bob.learn.em.JFABase
A JFABase object to be compared.r_epsilon : float
Relative precision.a_epsilon : float
Absolute precision.Returns:
output : bool
True if it is similar, otherwise false.
- load(hdf5) → None¶
Load the configuration of the JFABase to a given HDF5 file
Parameters:
hdf5 : bob.io.base.HDF5File
An HDF5 file open for reading
- resize(rU, rV) → None¶
Resets the dimensionality of the subspace U and V. U and V are hence uninitialized
Parameters:
rU : int
Size of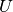 (Within client variation matrix)
(Within client variation matrix)rV : int
Size of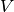 (Between client variation matrix)
(Between client variation matrix)
- save(hdf5) → None¶
Save the configuration of the JFABase to a given HDF5 file
Parameters:
hdf5 : bob.io.base.HDF5File
An HDF5 file open for writing
- shape¶
(int,int, int, int) <– A tuple that represents the number of gaussians, dimensionality of each Gaussian, dimensionality of the
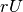 (within client variability matrix) and dimensionality of the
(within client variability matrix) and dimensionality of the
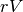 (between client variability matrix) (#Gaussians, #Inputs,
#rU, #rV).
(between client variability matrix) (#Gaussians, #Inputs,
#rU, #rV).
- supervector_length¶
int <– Returns the supervector length.NGaussians x NInputs: Number of Gaussian components by the feature dimensionality
@warning An exception is thrown if no Universal Background Model has been set yet.
- u¶
array_like <float, 2D> <– Returns the
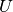 matrix (within
client variability matrix)
matrix (within
client variability matrix)
- ubm¶
bob.learn.em.GMMMachine <– Returns the UBM (Universal Background Model
- v¶
array_like <float, 2D> <– Returns the
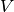 matrix (between
client variability matrix)
matrix (between
client variability matrix)
- class bob.learn.em.JFAMachine¶
Bases: object
A JFAMachine. An attached bob.learn.em.JFABase should be provided for Joint Factor Analysis. The bob.learn.em.JFAMachine carries information about the speaker factors
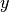 and
and  , whereas a
bob.learn.em.JFABase carries information about the matrices
, whereas a
bob.learn.em.JFABase carries information about the matrices
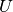 ,
, 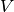 and
and 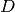 .
.References: [Vogt2008] [McCool2013]
Constructor Documentation:
- bob.learn.em.JFAMachine (jfa_base)
- bob.learn.em.JFAMachine (other)
- bob.learn.em.JFAMachine (hdf5)
Constructor. Builds a new JFAMachine
Parameters:
jfa_base : bob.learn.em.JFABase
The JFABase associated with this machineother : bob.learn.em.JFAMachine
A JFAMachine object to be copied.hdf5 : bob.io.base.HDF5File
An HDF5 file open for readingClass Members:
- estimate_ux(stats, input) → None¶
Estimates Ux (LPT assumption) given GMM statistics.
Estimates Ux from the GMM statistics considering the LPT assumption, that is the latent session variable x is approximated using the UBM.
Parameters:
stats : bob.learn.em.GMMStats
Statistics of the GMMinput : array_like <float, 1D>
Input vector
- estimate_x(stats, input) → None¶
Estimates the session offset x (LPT assumption) given GMM statistics.
Estimates x from the GMM statistics considering the LPT assumption, that is the latent session variable x is approximated using the UBM
Parameters:
stats : bob.learn.em.GMMStats
Statistics of the GMMinput : array_like <float, 1D>
Input vector
- forward_ux(stats, ux) → None¶
Computes a score for the given UBM statistics and given the Ux vector
Parameters:
stats : bob.learn.em.GMMStats
Statistics as inputux : array_like <float, 1D>
Input vector
- is_similar_to(other[, r_epsilon][, a_epsilon]) → output¶
Compares this JFAMachine with the other one to be approximately the same.
The optional values r_epsilon and a_epsilon refer to the relative and absolute precision for the weights, biases and any other values internal to this machine.
Parameters:
other : bob.learn.em.JFAMachine
A JFAMachine object to be compared.r_epsilon : float
Relative precision.a_epsilon : float
Absolute precision.Returns:
output : bool
True if it is similar, otherwise false.
- jfa_base¶
bob.learn.em.JFABase <– The JFABase attached to this machine
- load(hdf5) → None¶
Load the configuration of the JFAMachine to a given HDF5 file
Parameters:
hdf5 : bob.io.base.HDF5File
An HDF5 file open for reading
- log_likelihood(stats) → None¶
Computes the log-likelihood of the given samples
Note
the __call__() function is an alias for this function.
Parameters:
stats : bob.learn.em.GMMStats
Statistics as input
- save(hdf5) → None¶
Save the configuration of the JFAMachine to a given HDF5 file
Parameters:
hdf5 : bob.io.base.HDF5File
An HDF5 file open for writing
- shape¶
(int,int, int, int) <– A tuple that represents the number of gaussians, dimensionality of each Gaussian, dimensionality of the rU (within client variability matrix) and dimensionality of the rV (between client variability matrix) (#Gaussians, #Inputs, #rU, #rV).
- supervector_length¶
int <– Returns the supervector length.NGaussians x NInputs: Number of Gaussian components by the feature dimensionality
@warning An exception is thrown if no Universal Background Model has been set yet.
- x¶
array_like <float, 1D> <– Returns the
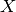 session factor. Eq
(29) from [McCool2013]
session factor. Eq
(29) from [McCool2013]The latent variable
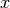 (last one computed). This is a feature
provided for convenience, but this attribute is not ‘part’ of the
machine. The session latent variable
(last one computed). This is a feature
provided for convenience, but this attribute is not ‘part’ of the
machine. The session latent variable 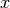 is indeed not
class-specific, but depends on the sample considered. Furthermore, it
is not saved into the machine or used when comparing machines.
is indeed not
class-specific, but depends on the sample considered. Furthermore, it
is not saved into the machine or used when comparing machines.
- y¶
array_like <float, 1D> <– Returns the
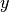 speaker factor. Eq
(30) from [McCool2013]
speaker factor. Eq
(30) from [McCool2013]
- z¶
array_like <float, 1D> <– Returns the
 speaker factor. Eq
(31) from [McCool2013]
speaker factor. Eq
(31) from [McCool2013]
- class bob.learn.em.JFATrainer¶
Bases: object
JFATrainerReferences: [Vogt2008,McCool2013]
Constructor Documentation:
- bob.learn.em.JFATrainer (other)
- bob.learn.em.JFATrainer ()
Constructor. Builds a new JFATrainer
Parameters:
other : bob.learn.em.JFATrainer
A JFATrainer object to be copied.Class Members:
- acc_d_a1¶
array_like <float, 1D> <– Accumulator updated during the E-step
- acc_d_a2¶
array_like <float, 1D> <– Accumulator updated during the E-step
- acc_u_a1¶
array_like <float, 3D> <– Accumulator updated during the E-step
- acc_u_a2¶
array_like <float, 2D> <– Accumulator updated during the E-step
- acc_v_a1¶
array_like <float, 3D> <– Accumulator updated during the E-step
- acc_v_a2¶
array_like <float, 2D> <– Accumulator updated during the E-step
- e_step_d(jfa_base, stats) → None¶
Call the 3rd e-step procedure (for the d subspace).
Parameters:
jfa_base : bob.learn.em.JFABase
JFABase Objectstats : bob.learn.em.GMMStats
GMMStats Object
- e_step_u(jfa_base, stats) → None¶
Call the 2nd e-step procedure (for the U subspace).
Parameters:
jfa_base : bob.learn.em.JFABase
JFABase Objectstats : bob.learn.em.GMMStats
GMMStats Object
- e_step_v(jfa_base, stats) → None¶
Call the 1st e-step procedure (for the V subspace).
Parameters:
jfa_base : bob.learn.em.JFABase
JFABase Objectstats : bob.learn.em.GMMStats
GMMStats Object
- enroll(jfa_machine, features, n_iter) → None¶
Parameters:
jfa_machine : bob.learn.em.JFAMachine
JFAMachine Objectfeatures : [bob.learn.em.GMMStats]
n_iter : int
Number of iterations
- finalize_d(jfa_base, stats) → None¶
Call the 3rd finalize procedure (for the d subspace).
Parameters:
jfa_base : bob.learn.em.JFABase
JFABase Objectstats : bob.learn.em.GMMStats
GMMStats Object
- finalize_u(jfa_base, stats) → None¶
Call the 2nd finalize procedure (for the U subspace).
Parameters:
jfa_base : bob.learn.em.JFABase
JFABase Objectstats : bob.learn.em.GMMStats
GMMStats Object
- finalize_v(jfa_base, stats) → None¶
Call the 1st finalize procedure (for the V subspace).
Parameters:
jfa_base : bob.learn.em.JFABase
JFABase Objectstats : bob.learn.em.GMMStats
GMMStats Object
- initialize(jfa_base, stats, rng) → None¶
Initialization before the EM steps
Parameters:
jfa_base : bob.learn.em.JFABase
JFABase Objectstats : bob.learn.em.GMMStats
GMMStats Objectrng : bob.core.random.mt19937
The Mersenne Twister mt19937 random generator used for the initialization of subspaces/arrays before the EM loop.
- m_step_d(jfa_base, stats) → None¶
Call the 3rd m-step procedure (for the d subspace).
Parameters:
jfa_base : bob.learn.em.JFABase
JFABase Objectstats : bob.learn.em.GMMStats
GMMStats Object
- m_step_u(jfa_base, stats) → None¶
Call the 2nd m-step procedure (for the U subspace).
Parameters:
jfa_base : bob.learn.em.JFABase
JFABase Objectstats : bob.learn.em.GMMStats
GMMStats Object
- m_step_v(jfa_base, stats) → None¶
Call the 1st m-step procedure (for the V subspace).
Parameters:
jfa_base : bob.learn.em.JFABase
JFABase Objectstats : bob.learn.em.GMMStats
GMMStats Object
- class bob.learn.em.KMeansMachine¶
Bases: object
This class implements a k-means classifier. See Section 9.1 of Bishop, “Pattern recognition and machine learning”, 2006
Constructor Documentation:
- bob.learn.em.KMeansMachine (n_means,n_inputs)
- bob.learn.em.KMeansMachine (other)
- bob.learn.em.KMeansMachine (hdf5)
- bob.learn.em.KMeansMachine ()
Creates a KMeansMachine
Parameters:
n_means : int
Number of meansn_inputs : int
Dimension of the feature vectorother : bob.learn.em.KMeansMachine
A KMeansMachine object to be copied.hdf5 : bob.io.base.HDF5File
An HDF5 file open for readingClass Members:
- get_closest_mean(input) → output¶
Calculate the index of the mean that is closest (in terms of square Euclidean distance) to the data sample, x.
Parameters:
input : array_like <float, 1D>
The data sample (feature vector)Returns:
output : (int, int)
Tuple containing the closest mean and the minimum distance from the input
- get_distance_from_mean(input, i) → output¶
Return the power of two of the square Euclidean distance of the sample, x, to the i’th mean.
Note
An exception is thrown if i is out of range.
Parameters:
input : array_like <float, 1D>
The data sample (feature vector)i : int
The index of the meanReturns:
output : float
Square Euclidean distance of the sample, x, to the i’th mean
- get_mean(i) → mean¶
Get the i’th mean.
Note
An exception is thrown if i is out of range.
Parameters:
i : int
Index of the meanReturns:
mean : array_like <float, 1D>
Mean array
- get_min_distance(input) → output¶
Output the minimum (Square Euclidean) distance between the input and the closest mean
Parameters:
input : array_like <float, 1D>
The data sample (feature vector)Returns:
output : float
The minimum distance
- get_variances_and_weights_for_each_cluster(input) → output¶
For each mean, find the subset of the samples that is closest to that mean, and calculate 1) the variance of that subset (the cluster variance) 2) the proportion of the samples represented by that subset (the cluster weight)
Parameters:
input : array_like <float, 2D>
The data sample (feature vector)Returns:
output : (array_like <float, 2D>, array_like <float, 1D>)
A tuple with the variances and the weights respectively
- is_similar_to(other[, r_epsilon][, a_epsilon]) → output¶
Compares this KMeansMachine with the other one to be approximately the same.
The optional values r_epsilon and a_epsilon refer to the relative and absolute precision for the weights, biases and any other values internal to this machine.
Parameters:
other : bob.learn.em.KMeansMachine
A KMeansMachine object to be compared.r_epsilon : float
Relative precision.a_epsilon : float
Absolute precision.Returns:
output : bool
True if it is similar, otherwise false.
- load(hdf5) → None¶
Load the configuration of the KMeansMachine to a given HDF5 file
Parameters:
hdf5 : bob.io.base.HDF5File
An HDF5 file open for reading
- means¶
array_like <float, 2D> <– The means
- resize(n_means, n_inputs) → None¶
Allocates space for the statistics and resets to zero.
Parameters:
n_means : int
Number of meansn_inputs : int
Dimensionality of the feature vector
- save(hdf5) → None¶
Save the configuration of the KMeansMachine to a given HDF5 file
Parameters:
hdf5 : bob.io.base.HDF5File
An HDF5 file open for writing
- set_mean(i, mean) → None¶
Set the i’th mean.
Note
An exception is thrown if i is out of range.
Parameters:
i : int
Index of the meanmean : array_like <float, 1D>
Mean array
- shape¶
(int,int) <– A tuple that represents the number of means and dimensionality of the feature vector``(n_means, dim)``.
- class bob.learn.em.KMeansTrainer¶
Bases: object
Trains a KMeans machine.This class implements the expectation-maximization algorithm for a k-means machine.See Section 9.1 of Bishop, “Pattern recognition and machine learning”, 2006It uses a random initialization of the means followed by the expectation-maximization algorithm
Constructor Documentation:
- bob.learn.em.KMeansTrainer (initialization_method)
- bob.learn.em.KMeansTrainer (other)
- bob.learn.em.KMeansTrainer ()
Creates a KMeansTrainer
Parameters:
initialization_method : str
The initialization method of the means. Possible values are: ‘RANDOM’, ‘RANDOM_NO_DUPLICATE’, ‘KMEANS_PLUS_PLUS’other : bob.learn.em.KMeansTrainer
A KMeansTrainer object to be copied.Class Members:
- average_min_distance¶
str <– Average min (square Euclidean) distance. Useful to parallelize the E-step.
- compute_likelihood(kmeans_machine) → None¶
This functions returns the average min (Square Euclidean) distance (average distance to the closest mean)
Parameters:
kmeans_machine : bob.learn.em.KMeansMachine
KMeansMachine Object
- e_step(kmeans_machine, data) → None¶
Compute the E-step, which is basically the distances
Accumulate across the dataset: -zeroeth and first order statistics -average (Square Euclidean) distance from the closest mean
Parameters:
kmeans_machine : bob.learn.em.KMeansMachine
KMeansMachine Objectdata : array_like <float, 2D>
Input data
- first_order_statistics¶
array_like <float, 2D> <– Returns the internal statistics. Useful to parallelize the E-step
- initialization_method¶
str <– Initialization method.
- Possible values:
RANDOM: Random initialization
RANDOM_NO_DUPLICATE: Random initialization without repetition
KMEANS_PLUS_PLUS: Apply the kmeans++ initialization http://en.wikipedia.org/wiki/K-means%2B%2B
- initialize(kmeans_machine, data, rng) → None¶
Initialise the means randomly
Data is split into as many chunks as there are means, then each mean is set to a random example within each chunk.
Parameters:
kmeans_machine : bob.learn.em.KMeansMachine
KMeansMachine Objectdata : array_like <float, 2D>
Input datarng : bob.core.random.mt19937
The Mersenne Twister mt19937 random generator used for the initialization of subspaces/arrays before the EM loop.
- m_step(kmeans_machine, data) → None¶
Updates the mean based on the statistics from the E-step
Parameters:
kmeans_machine : bob.learn.em.KMeansMachine
KMeansMachine Objectdata : array_like <float, 2D>
Ignored.
- reset_accumulators(kmeans_machine) → None¶
Reset the statistics accumulators to the correct size and a value of zero.
Parameters:
kmeans_machine : bob.learn.em.KMeansMachine
KMeansMachine Object
- zeroeth_order_statistics¶
array_like <float, 1D> <– Returns the internal statistics. Useful to parallelize the E-step
- class bob.learn.em.MAP_GMMTrainer¶
Bases: object
This class implements the maximum a posteriori M-step of the expectation-maximization algorithm for a GMM Machine. The prior parameters are encoded in the form of a GMM (e.g. a universal background model). The EM algorithm thus performs GMM adaptation.
Constructor Documentation:
- bob.learn.em.MAP_GMMTrainer (prior_gmm, relevance_factor, [update_means], [update_variances], [update_weights], [mean_var_update_responsibilities_threshold])
- bob.learn.em.MAP_GMMTrainer (prior_gmm, alpha, [update_means], [update_variances], [update_weights], [mean_var_update_responsibilities_threshold])
- bob.learn.em.MAP_GMMTrainer (other)
Creates a MAP_GMMTrainer
Additionally to the copy constructor, there are two different ways to call this constructor, one using the relevance_factor and one using the alpha, both which have the same signature. Hence, the only way to differentiate the two functions is by using keyword arguments.
Parameters:
prior_gmm : bob.learn.em.GMMMachine
The prior GMM to be adapted (Universal Background Model UBM).relevance_factor : float
If set the Reynolds Adaptation procedure will be applied. See Eq (14) from [Reynolds2000]alpha : float
Set directly the alpha parameter (Eq (14) from [Reynolds2000]), ignoring zeroth order statistics as a weighting factor.update_means : bool
[Default: True] Update means on each iterationupdate_variances : bool
[Default: True] Update variances on each iterationupdate_weights : bool
[Default: True] Update weights on each iterationmean_var_update_responsibilities_threshold : float
[Default: min_float] Threshold over the responsibilities of the Gaussians Equations 9.24, 9.25 of Bishop, Pattern recognition and machine learning, 2006 require a division by the responsibilities, which might be equal to zero because of numerical issue. This threshold is used to avoid such divisions.other : bob.learn.em.MAP_GMMTrainer
A MAP_GMMTrainer object to be copied.Class Members:
- alpha¶
float <– Set directly the alpha parameter (Eq (14) from [Reynolds2000]), ignoring zeroth order statistics as a weighting factor.
- compute_likelihood(gmm_machine) → None¶
This functions returns the average min (Square Euclidean) distance (average distance to the closest mean)
Parameters:
gmm_machine : bob.learn.em.GMMMachine
GMMMachine Object
- e_step(gmm_machine, data) → None¶
Calculates and saves statistics across the dataset and saves these as gmm_statistics.
Calculates the average log likelihood of the observations given the GMM,and returns this in average_log_likelihood.The statistics, gmm_statistics, will be used in the m_step() that follows.
Parameters:
gmm_machine : bob.learn.em.GMMMachine
GMMMachine Objectdata : array_like <float, 2D>
Input data
- gmm_statistics¶
GMMStats <– The GMM statistics that were used internally in the E- and M-steps
Setting and getting the internal GMM statistics might be useful to parallelize the GMM training.
- initialize(gmm_machine, data) → None¶
Initialization before the EM steps
Parameters:
gmm_machine : bob.learn.em.GMMMachine
GMMMachine Objectdata : array_like <float, 2D>
Ignored.
- m_step(gmm_machine, data) → None¶
Performs a maximum a posteriori (MAP) update of the GMM parameters using the accumulated statistics in gmm_statistics and the parameters of the prior model
Parameters:
gmm_machine : bob.learn.em.GMMMachine
GMMMachine Objectdata : array_like <float, 2D>
Ignored.
- relevance_factor¶
float <– If set the reynolds_adaptation parameters, will apply the Reynolds Adaptation Factor. See Eq (14) from [Reynolds2000]
- class bob.learn.em.ML_GMMTrainer¶
Bases: object
This class implements the maximum likelihood M-step of the expectation-maximisation algorithm for a GMM Machine.
Constructor Documentation:
- bob.learn.em.ML_GMMTrainer (update_means, [update_variances], [update_weights], [mean_var_update_responsibilities_threshold])
- bob.learn.em.ML_GMMTrainer (other)
Creates a ML_GMMTrainer
Parameters:
update_means : bool
Update means on each iterationupdate_variances : bool
Update variances on each iterationupdate_weights : bool
Update weights on each iterationmean_var_update_responsibilities_threshold : float
Threshold over the responsibilities of the Gaussians Equations 9.24, 9.25 of Bishop, Pattern recognition and machine learning, 2006 require a division by the responsibilities, which might be equal to zero because of numerical issue. This threshold is used to avoid such divisions.other : bob.learn.em.ML_GMMTrainer
A ML_GMMTrainer object to be copied.Class Members:
- compute_likelihood(gmm_machine) → None¶
This functions returns the average min (Square Euclidean) distance (average distance to the closest mean)
Parameters:
gmm_machine : bob.learn.em.GMMMachine
GMMMachine Object
- e_step(gmm_machine, data) → None¶
Calculates and saves statistics across the dataset,and saves these as m_ss.
Calculates the average log likelihood of the observations given the GMM,and returns this in average_log_likelihood.The statistics, gmm_statistics, will be used in the m_step() that follows.
Parameters:
gmm_machine : bob.learn.em.GMMMachine
GMMMachine Objectdata : array_like <float, 2D>
Input data
- gmm_statistics¶
GMMStats <– The GMM statistics that were used internally in the E- and M-steps
Setting and getting the internal GMM statistics might be useful to parallelize the GMM training.
- initialize(gmm_machine, data) → None¶
Initialization before the EM steps
Parameters:
gmm_machine : bob.learn.em.GMMMachine
GMMMachine Objectdata : array_like <float, 2D>
Ignored.
- m_step(gmm_machine, data) → None¶
Performs a maximum likelihood (ML) update of the GMM parameters using the accumulated statistics in gmm_statistics
See Section 9.2.2 of Bishop, “Pattern recognition and machine learning”, 2006
Parameters:
gmm_machine : bob.learn.em.GMMMachine
GMMMachine Objectdata : array_like <float, 2D>
Ignored.
- class bob.learn.em.PLDABase¶
Bases: object
This class is a container for the
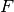 (between class variantion
matrix),
(between class variantion
matrix), 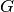 (within class variantion matrix) and
(within class variantion matrix) and 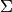 matrices and the mean vector
matrices and the mean vector 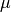 of a PLDA model. This
alsoprecomputes useful matrices to make the model scalable.References:
[ElShafey2014,PrinceElder2007,LiFu2012]
of a PLDA model. This
alsoprecomputes useful matrices to make the model scalable.References:
[ElShafey2014,PrinceElder2007,LiFu2012]Constructor Documentation:
- bob.learn.em.PLDABase (dim_d,dim_f,dim_g,variance_threshold)
- bob.learn.em.PLDABase (other)
- bob.learn.em.PLDABase (hdf5)
Constructor, builds a new PLDABase.
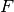 ,
, 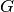 and
and
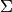 are initialized to the ‘eye’ matrix (matrix with 1’s
on the diagonal and 0 outside), and
are initialized to the ‘eye’ matrix (matrix with 1’s
on the diagonal and 0 outside), and 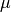 is initialized to
0.
is initialized to
0.Parameters:
dim_d : int
Dimensionality of the feature vector.dim_f : int
Size of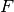 (between class variantion matrix).
(between class variantion matrix).dim_g : int
Size of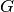 (within class variantion matrix).
(within class variantion matrix).variance_threshold : float
The smallest possible value of the variance (Ignored if set to 0.)other : bob.learn.em.PLDABase
A PLDABase object to be copied.hdf5 : bob.io.base.HDF5File
An HDF5 file open for readingClass Members:
- clear_maps() → None¶
Clears the maps (
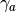 and loglike_constterm_a).
and loglike_constterm_a).
- compute_gamma(a, res) → None¶
Tells if the
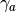 matrix for a given a (number of
samples) exists.
matrix for a given a (number of
samples) exists. 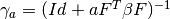
Parameters:
a : int
Indexres : array_like <float, 2D>
Input data
- compute_log_like_const_term(a, res) → None¶
Computes the log likelihood constant term for a given
 (number of samples), given the provided
(number of samples), given the provided 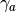 matrix.
matrix.
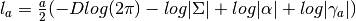
Parameters:
a : int
Indexres : array_like <float, 2D>
Input data
- compute_log_likelihood_point_estimate(xij, hi, wij) → output¶
Gets the log-likelihood of an observation, given the current model and the latent variables (point estimate).This will basically compute
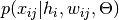 , given by
, given by
![\mathcal{N}(x_{ij}|[\mu + F h_{i} + G w_{ij} +
\epsilon_{ij}, \Sigma])](_images/math/5e92c5e2291f42039327a6b6b5af6d62099365f4.png) , which is in logarithm,
, which is in logarithm, 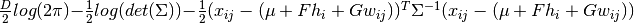
Parameters:
xij : array_like <float, 1D>
hi : array_like <float, 1D>
wij : array_like <float, 1D>
Returns:
output : float
- f¶
array_like <float, 2D> <– Returns the
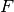 matrix (between
class variantion matrix)
matrix (between
class variantion matrix)
- g¶
array_like <float, 2D> <– Returns the
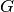 matrix (between
class variantion matrix)
matrix (between
class variantion matrix)
- get_add_gamma(a) → output¶
Gets the
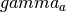 matrix for a given
matrix for a given 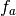 (number of
samples).
(number of
samples). 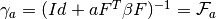 .Tries to find it from the base machine and then
from this machine.
.Tries to find it from the base machine and then
from this machine.Parameters:
a : int
IndexReturns:
output : array_like <float, 2D>
- get_add_log_like_const_term(a) → output¶
Gets the log likelihood constant term for a given
 (number
of samples).
(number
of samples). 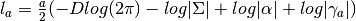
Parameters:
a : int
IndexReturns:
output : float
- get_gamma(a) → output¶
Gets the
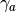 matrix for a given
matrix for a given  (number of
samples).
(number of
samples). 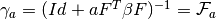
Parameters:
a : int
IndexReturns:
output : array_like <float, 2D>
Get the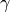 matrix
matrix
- get_log_like_const_term(a) → output¶
Gets the log likelihood constant term for a given
 (number
of samples).
(number
of samples). 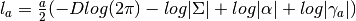
Parameters:
a : int
IndexReturns:
output : float
- has_gamma(a) → output¶
Tells if the
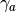 matrix for a given a (number of
samples) exists.
matrix for a given a (number of
samples) exists. 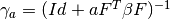
Parameters:
a : int
IndexReturns:
output : bool
- has_log_like_const_term(a) → output¶
Tells if the log likelihood constant term for a given
 (number of samples) exists in this machine (does not check the base
machine).
(number of samples) exists in this machine (does not check the base
machine). 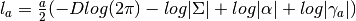
Parameters:
a : int
IndexReturns:
output : bool
- is_similar_to(other[, r_epsilon][, a_epsilon]) → output¶
Compares this PLDABase with the other one to be approximately the same.
The optional values r_epsilon and a_epsilon refer to the relative and absolute precision for the weights, biases and any other values internal to this machine.
Parameters:
other : bob.learn.em.PLDABase
A PLDABase object to be compared.r_epsilon : float
Relative precision.a_epsilon : float
Absolute precision.Returns:
output : bool
True if it is similar, otherwise false.
- load(hdf5) → None¶
Load the configuration of the PLDABase to a given HDF5 file
Parameters:
hdf5 : bob.io.base.HDF5File
An HDF5 file open for reading
- mu¶
array_like <float, 1D> <– Gets the
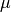 mean vector of the
PLDA model
mean vector of the
PLDA model
- resize(dim_d, dim_f, dim_g) → None¶
Resizes the dimensionality of the PLDA model. Paramaters
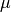 ,
, 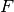 ,
, 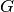 and
and 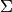 are
reinitialized.
are
reinitialized.Parameters:
dim_d : int
Dimensionality of the feature vector.dim_f : int
Size of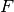 (between class variantion matrix).
(between class variantion matrix).dim_g : int
Size of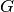 (within class variantion matrix).
(within class variantion matrix).
- save(hdf5) → None¶
Save the configuration of the PLDABase to a given HDF5 file
Parameters:
hdf5 : bob.io.base.HDF5File
An HDF5 file open for writing
- shape¶
(int,int, int) <– A tuple that represents the dimensionality of the feature vector dim_d, the
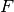 matrix and the
matrix and the 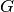 matrix.
matrix.
- sigma¶
array_like <float, 1D> <– Gets the
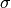 (diagonal)
covariance matrix of the PLDA model
(diagonal)
covariance matrix of the PLDA model
- variance_threshold¶
float <–
- class bob.learn.em.PLDAMachine¶
Bases: object
This class is a container for an enrolled identity/class. It contains information extracted from the enrollment samples. It should be used in combination with a PLDABase instance.
References: [ElShafey2014], [PrinceElder2007], [LiFu2012]
Constructor Documentation:
- bob.learn.em.PLDAMachine (plda_base)
- bob.learn.em.PLDAMachine (other)
- bob.learn.em.PLDAMachine (hdf5,plda_base)
Constructor, builds a new PLDAMachine.
Parameters:
plda_base : bob.learn.em.PLDABase
other : bob.learn.em.PLDAMachine
A PLDAMachine object to be copied.hdf5 : bob.io.base.HDF5File
An HDF5 file open for readingClass Members:
- clear_maps() → None¶
Clears the maps (
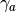 and loglike_constterm_a).
and loglike_constterm_a).
- compute_log_likelihood(sample, with_enrolled_samples) → output¶
Compute the log-likelihood of the given sample and (optionally) the enrolled samples
Parameters:
sample : array_like <float, 1D>,array_like <float, 2D>
Samplewith_enrolled_samples : bool
Returns:
output : float
The log-likelihood
- get_add_gamma(a) → output¶
Gets the
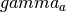 matrix for a given
matrix for a given 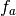 (number of
samples).
(number of
samples). 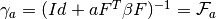 .Tries to find it from the base machine and then
from this machine.
.Tries to find it from the base machine and then
from this machine.Parameters:
a : int
IndexReturns:
output : array_like <float, 2D>
- get_add_log_like_const_term(a) → output¶
Gets the log likelihood constant term for a given
 (number
of samples).
(number
of samples). 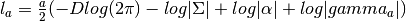
Parameters:
a : int
IndexReturns:
output : float
- get_gamma(a) → output¶
Gets the
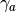 matrix for a given
matrix for a given  (number of
samples).
(number of
samples). 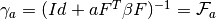
Parameters:
a : int
IndexReturns:
output : array_like <float, 2D>
Get the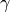 matrix
matrix
- get_log_like_const_term(a) → output¶
Gets the log likelihood constant term for a given
 (number
of samples).
(number
of samples). 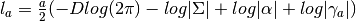
Parameters:
a : int
IndexReturns:
output : float
- has_gamma(a) → output¶
Tells if the
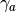 matrix for a given a (number of
samples) exists.
matrix for a given a (number of
samples) exists. 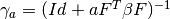
Parameters:
a : int
IndexReturns:
output : bool
- has_log_like_const_term(a) → output¶
Tells if the log likelihood constant term for a given
 (number of samples) exists in this machine (does not check the base
machine).
(number of samples) exists in this machine (does not check the base
machine). 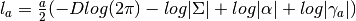
Parameters:
a : int
IndexReturns:
output : bool
- is_similar_to(other[, r_epsilon][, a_epsilon]) → output¶
Compares this PLDAMachine with the other one to be approximately the same.
The optional values r_epsilon and a_epsilon refer to the relative and absolute precision for the weights, biases and any other values internal to this machine.
Parameters:
other : bob.learn.em.PLDAMachine
A PLDAMachine object to be compared.r_epsilon : float
Relative precision.a_epsilon : float
Absolute precision.Returns:
output : bool
True if it is similar, otherwise false.
- load(hdf5) → None¶
Load the configuration of the PLDAMachine to a given HDF5 file
Parameters:
hdf5 : bob.io.base.HDF5File
An HDF5 file open for reading
- log_likelihood¶
float <–
- log_likelihood_ratio(samples) → output¶
Computes a log likelihood ratio from a 1D or 2D blitz::Array
Parameters:
samples : array_like <float, 1D>,array_like <float, 2D>
SampleReturns:
output : float
The log-likelihood ratio
- n_samples¶
int <– Number of enrolled samples
- plda_base¶
bob.learn.em.PLDABase <– The PLDABase attached to this machine
- save(hdf5) → None¶
Save the configuration of the PLDAMachine to a given HDF5 file
Parameters:
hdf5 : bob.io.base.HDF5File
An HDF5 file open for writing
- shape¶
(int,int, int) <– A tuple that represents the dimensionality of the feature vector dim_d, the
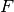 matrix and the
matrix and the 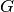 matrix.
matrix.
- w_sum_xit_beta_xi¶
float <– Gets the
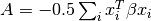 value
value
- weighted_sum¶
array_like <float, 1D> <– Get/Set
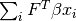 value
value
- class bob.learn.em.PLDATrainer¶
Bases: object
This class can be used to train the
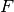 ,
, 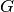 and
and
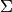 matrices and the mean vector
matrices and the mean vector 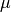 of a PLDA
model.References: [ElShafey2014],[PrinceElder2007]_,[LiFu2012]_
of a PLDA
model.References: [ElShafey2014],[PrinceElder2007]_,[LiFu2012]_Constructor Documentation:
- bob.learn.em.PLDATrainer (use_sum_second_order)
- bob.learn.em.PLDATrainer (other)
- bob.learn.em.PLDATrainer ()
- Default constructor.
- Initializes a new PLDA trainer. The training stage will place the resulting components in the PLDABase.
Parameters:
other : bob.learn.em.PLDATrainer
A PLDATrainer object to be copied.use_sum_second_order : bool
Class Members:
- e_step(plda_base, data) → None¶
Expectation step before the EM steps
Parameters:
plda_base : bob.learn.em.PLDABase
PLDAMachine Objectdata : list
- enroll(plda_machine, data) → None¶
Main procedure for enrolling a PLDAMachine
Parameters:
plda_machine : bob.learn.em.PLDAMachine
PLDAMachine Objectdata : list
- finalize(plda_base, data) → None¶
finalize before the EM steps
Parameters:
plda_base : bob.learn.em.PLDABase
PLDAMachine Objectdata : list
- init_f_method¶
str <– The method used for the initialization of
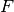 .
.Possible values are: (‘RANDOM_F’, ‘BETWEEN_SCATTER’)
- init_g_method¶
str <– The method used for the initialization of
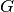 .
.Possible values are: (‘RANDOM_G’, ‘WITHIN_SCATTER’)
- init_sigma_method¶
str <– The method used for the initialization of
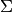 .
.Possible values are: (‘RANDOM_SIGMA’, ‘VARIANCE_G’, ‘CONSTANT’, ‘VARIANCE_DATA’)
- initialize(plda_base, data, rng) → None¶
Initialization before the EM steps
Parameters:
plda_base : bob.learn.em.PLDABase
PLDAMachine Objectdata : list
rng : bob.core.random.mt19937
The Mersenne Twister mt19937 random generator used for the initialization of subspaces/arrays before the EM loop.
- is_similar_to(other[, r_epsilon][, a_epsilon]) → output¶
Compares this PLDATrainer with the other one to be approximately the same.
The optional values r_epsilon and a_epsilon refer to the relative and absolute precision for the weights, biases and any other values internal to this machine.
Parameters:
other : bob.learn.em.PLDAMachine
A PLDAMachine object to be compared.r_epsilon : float
Relative precision.a_epsilon : float
Absolute precision.Returns:
output : bool
True if it is similar, otherwise false.
- m_step(plda_base, data) → None¶
Maximization step
Parameters:
plda_base : bob.learn.em.PLDABase
PLDAMachine Objectdata : list
- use_sum_second_order¶
bool <– Tells whether the second order statistics are stored during the training procedure, or only their sum.
- z_first_order¶
array_like <float, 2D> <–
- z_second_order¶
array_like <float, 3D> <–
- z_second_order_sum¶
array_like <float, 2D> <–
- bob.learn.em.linear_scoring(models, ubm, test_stats, test_channelOffset, frame_length_normalisation) → output¶
Parameters:
models : [bob.learn.em.GMMMachine]
ubm : bob.learn.em.GMMMachine
test_stats : [bob.learn.em.GMMStats]
test_channelOffset : [array_like<float,1>]
frame_length_normalisation : bool
Returns:
output : array_like<float,1>
Score
- bob.learn.em.tnorm(rawscores_probes_vs_models, rawscores_probes_vs_tmodels) → output¶
Normalise raw scores with T-Norm
Parameters:
rawscores_probes_vs_models : array_like <float, 2D>
Raw set of scoresrawscores_probes_vs_tmodels : array_like <float, 2D>
T-Scores (raw scores of the T probes against the models)Returns:
output : array_like <float, 2D>
The scores T Normalized
- bob.learn.em.train(trainer, machine, data, max_iterations=50, convergence_threshold=None, initialize=True, rng=None)¶
Trains a machine given a trainer and the proper data
- Parameters:
- trainer : one of KMeansTrainer, MAP_GMMTrainer, ML_GMMTrainer, ISVTrainer, IVectorTrainer, PLDATrainer, EMPCATrainer
- A trainer mechanism
- machine : one of KMeansMachine, GMMMachine, ISVBase, IVectorMachine, PLDAMachine, bob.learn.linear.Machine
- A container machine
- data : array_like <float, 2D>
- The data to be trained
- max_iterations : int
- The maximum number of iterations to train a machine
- convergence_threshold : float
- The convergence threshold to train a machine. If None, the training procedure will stop with the iterations criteria
- initialize : bool
- If True, runs the initialization procedure
- rng : bob.core.random.mt19937
- The Mersenne Twister mt19937 random generator used for the initialization of subspaces/arrays before the EM loop
- bob.learn.em.train_jfa(trainer, jfa_base, data, max_iterations=10, initialize=True, rng=None)¶
Trains a bob.learn.em.JFABase given a bob.learn.em.JFATrainer and the proper data
- Parameters:
- trainer : bob.learn.em.JFATrainer
- A JFA trainer mechanism
- jfa_base : bob.learn.em.JFABase
- A container machine
- data : [[bob.learn.em.GMMStats]]
- The data to be trained
- max_iterations : int
- The maximum number of iterations to train a machine
- initialize : bool
- If True, runs the initialization procedure
- rng : bob.core.random.mt19937
- The Mersenne Twister mt19937 random generator used for the initialization of subspaces/arrays before the EM loops
- bob.learn.em.znorm(rawscores_probes_vs_models, rawscores_zprobes_vs_models) → output¶
Normalise raw scores with Z-Norm
Parameters:
rawscores_probes_vs_models : array_like <float, 2D>
Raw set of scoresrawscores_zprobes_vs_models : array_like <float, 2D>
Z-Scores (raw scores of the Z probes against the models)Returns:
output : array_like <float, 2D>
The scores T Normalized
- bob.learn.em.ztnorm(rawscores_probes_vs_models, rawscores_zprobes_vs_models, rawscores_probes_vs_tmodels, rawscores_zprobes_vs_tmodels, mask_zprobes_vs_tmodels_istruetrial) → output¶
Normalise raw scores with ZT-Norm.Assume that znorm and tnorm have no common subject id.
Parameters:
rawscores_probes_vs_models : array_like <float, 2D>
Raw set of scoresrawscores_zprobes_vs_models : array_like <float, 2D>
Z-Scores (raw scores of the Z probes against the models)rawscores_probes_vs_tmodels : array_like <float, 2D>
T-Scores (raw scores of the T probes against the models)rawscores_zprobes_vs_tmodels : array_like <float, 2D>
ZT-Scores (raw scores of the Z probes against the T-models)mask_zprobes_vs_tmodels_istruetrial : array_like <float, 2D>
Returns:
output : array_like <float, 2D>
The scores ZT Normalized
